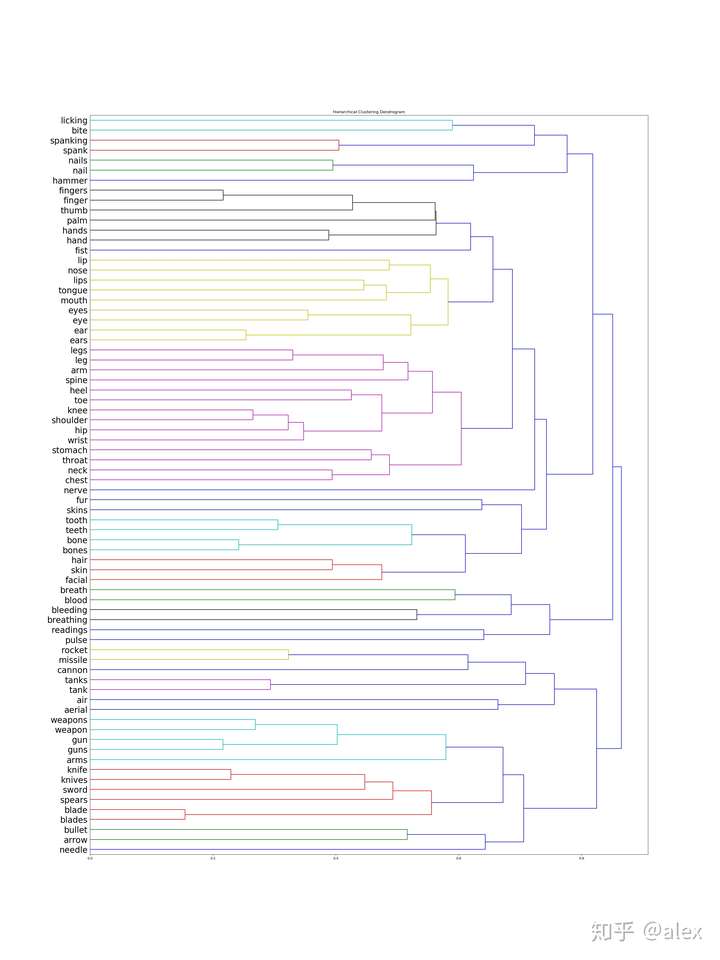
SCIPY CLUSTER CODE
You can see in the code I am using Agglomerative Clustering with 3 clusters, Euclidean distance parameters and ward as the linkage parameter. Sm.accuracy_score(target,HClustering.labels_)
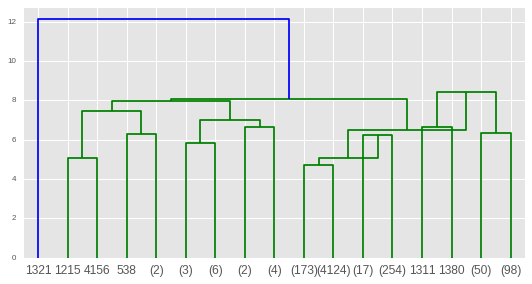
HClustering = AgglomerativeClustering(n_clusters=k, affinity="euclidean",linkage="ward") #based on the dendrogram we have two clusetes You will choose the method with the largest score. Doing this you will generate different accuracy score. In this step, you will generate a Hierarchical Cluster using the various affinity and linkage methods. Step 5: Generate the Hierarchical cluster. It means you should choose k=3, that is the number of clusters. And also the dataset has three types of species. Using the Iris dataset and its dendrogram, you can clearly see at distance approx y= 9 Line has divided into three clusters.

But at the last, you will take the distance metrics and linkage parameters on the accuracy score of the model. It’s just for the visualizing the dendrogram. In my case, I am using ward linkage parameter. There are three ways you can link the data points Ward, Complete and Average. Plt.title("Truncated Hierachial Clustering Dendrogram") z = linkage(data,"ward") #generate dendrogramĭendrogram(z,truncate_mode= "lastp", p =12, leaf_rotation=45,leaf_font_size=15, show_contracted=True) In this case, you will use the dendrogram. You should verify the number of clusters visually. In order to estimate the number of centroids. Step 4: Draw the Dendrogram of the dataset. Please note that you should always scale the data for accurate prediction. In this case, data is sepal length, sepal length, petal length, and petal width. Here data is the input variable(scaled data) and the target is the output variable. You can use your own dataset, but I am using only the default Iris dataset loaded from the Sklearn. If you want to read about it then here is the link for the t_printoptions. The first line np.set_printoptions(precision=4,suppress=True ) method will tell the python interpreter to use float datapoints up to 4 digits after the decimal. Np.set_printoptions(precision=4,suppress=True) Step 2: Import the libraries for the Data Visualization #Configure the output Here we are importing dendrogram, linkage, cluster, and cophenet from the packages. Easy Steps to Do Hierarchical Clustering in Python Step 1: Import the necessary Libraries for the Hierarchical Clustering import numpy as npįrom import dendrogram,linkageįrom import fclusterįrom import cophenetįrom sklearn.cluster import AgglomerativeClustering Popular Use Cases are Hospital Resource Management, Business Process Management, and Social Network Analysis. You will find many use cases for this type of clustering and some of them are DNA sequencing, Sentiment Analysis, Tracking Virus Diseases e.t.c. It allows you to see linkages, relatedness using the tree graph. In order to find the number of subgroups in the dataset, you use dendrogram. In this tutorial, I will use the popular approach Agglomerative way. There are two ways you can do Hierarchical clustering Agglomerative that is bottom-up approach clustering and Divisive uses top-down approaches for clustering. Each data point is linked to its nearest neighbors. Hierarchical Clustering uses the distance based approach between the neighbor datapoints for clustering.

